Automatic Data Extraction to Support Meta-Analysis Statistical Analysis
Overview
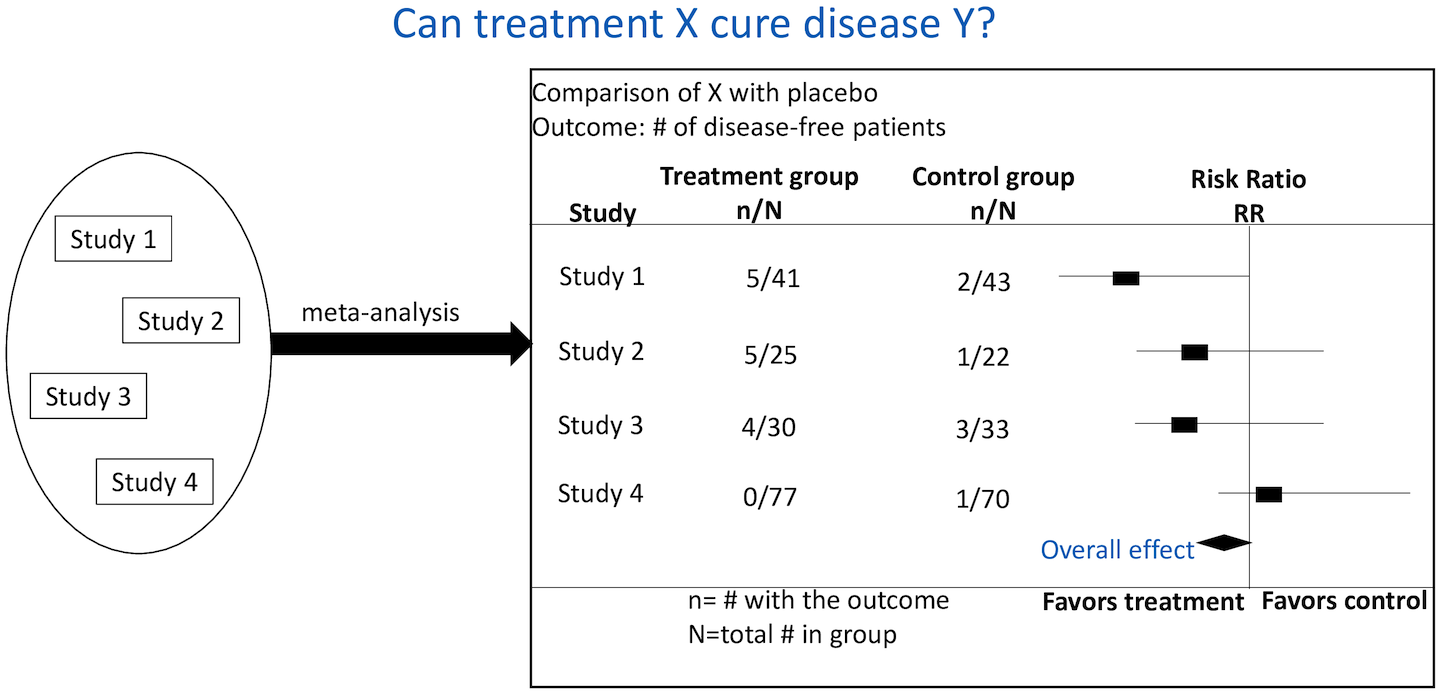
A meta-analysis is a statistical analysis that combines results of different studies that are focussed on the same disease or treatment to ascertain if a treatment is effective or not. For instance, if we want to determine if treatment X cures a certain disease/condition Y, we can (1) conduct a clinical trial, or (2) conduct a meta-analysis. Although clinical trials provide gold standard evidence, they are time consuming and expensive. Meta-analyses are therefore an alternative to clinical trials.
Meta-analysis involves extracting data from existing clinical trials reserch publications and combining the results to determine if the treatment is effective or not. However, the number of research publications is increasing rapidly and more articles are being published everyday. In this research, we propose a system that automatically extracts information from research publications. The system has the benefit of reducing the time taken in reading research articles to extract information.
Dataset
We created a corpus consisting of abstracts of breast cancer randomized controlled trials (RCTs) extracted from the PubMed database. We annotate the core components of clinical trials, i.e., Participants (P), Intervention (I), Control (C), and Outcome (O), commonly known as the PICO elements. For each component we annotate text snippets that describe them.
- Participants: we annotate the total number of participants, the number of participants in the intervention group, the number of participants in the control group, eligibility, age, condition, age, ethnicity, and location.
- Intervention and Control: we annotate the intervention and control treatments.
- Outcome: we annotate the outcomes that were measured, outcome measure, and number of participants that experienced an outcome.
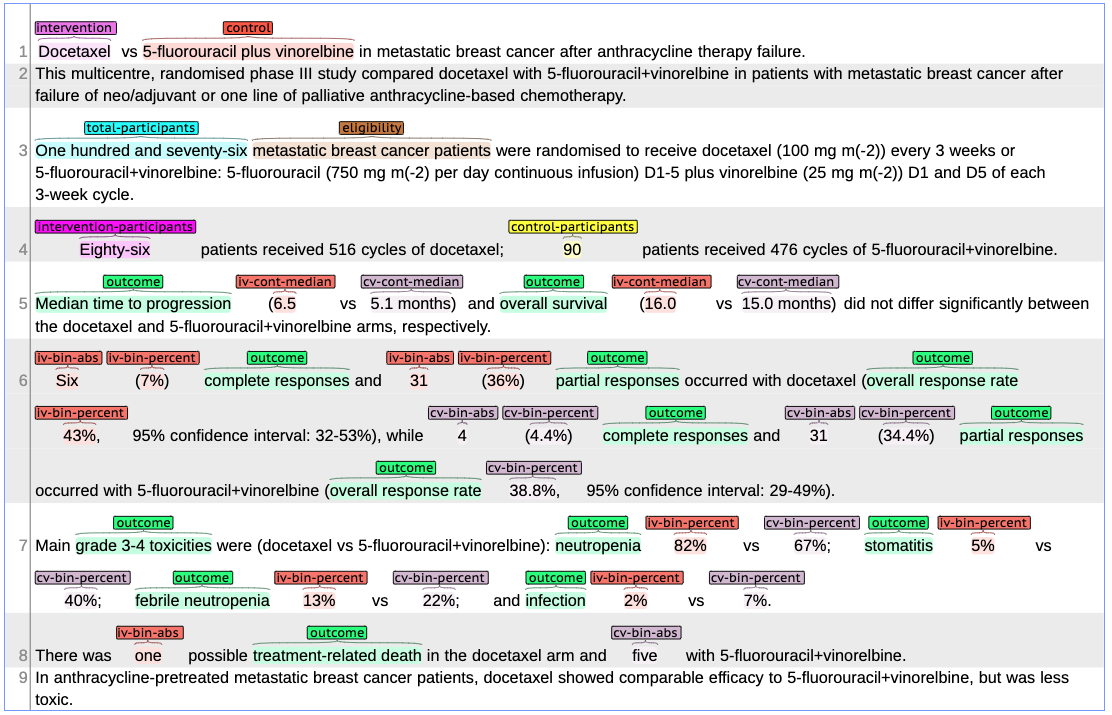
The abstracts were annotated using BRAT web annotation tool. Fig. 2 shows a sample abstract annotated in BRAT. The annotated data can be freely downloaded here. For detailed information on the annotation guidelines and entities please read our Paper.

Approach
Pre-processing
- Convert BRAT annotated texts to BIO format and save in csv format. The codes for converting from BRAT format to BIO can be found at this Github repository
- Acronym expansion. Research articles use acronyms to avoid repeating long terms and save space. In this research we identify acronyms by looking for terms in parenthesis that are between two and ten characters and use regular expresions to find expansion candidates in the sorrounding text. The code for acronym expansion can be found at Github repository.
Models
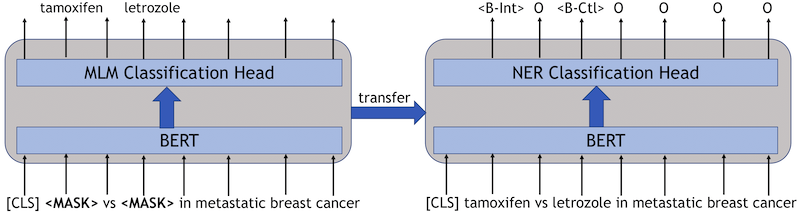
The information extraction is a named entity recognition (NER) task. We train three BERT-based NER models; BioBERT, BlueBERT, and Longformer. We use the standard BERT-based models for token classification as provided by huggingface. Since neural network models provide different results when initialized with different seeds, previous research suggests using different seeds and averaging the results. The code for token classification using BERT models can be found here.
Paper
Please read our paper for more details on this project.
Also check out our NER demo website and meta-analysis results visualization system.