Contextualized medication event extraction
Overview
This project outlines participation in the n2c2 2022 shared task (Track 1) on Contextalized Medication Event Extraction (CMED). The track consists of three subtasks:
- Named Entity Recognition (NER) - extracting medications/drugs mentions from clinical notes
- Event Classification - classifying the medication mentions into:
- Dispostion (there was medication change discussed)
- NoDisposition (there was no medication change discussed)
- Undetermined (unclear if a change was discussed).
- Context classification - classify the context of Disposition events into: (1) Action (start, stop), (2) Negation, (3) Temporality (past, present), (4) Certainty (hypothetical, conditional), and (5) Actor (patient, physician)
Here I will discuss our approach for the subtask 2 on Event classification.
Approach
Dataset
The data is not publicly available due to privacy and confidentiality reasons, but can be requested from the task organizers.
The dataset contains clinical notes annotated with BRAT web annotation tool. The organizers provided the brat annotated (.ann) files and the clinical notes as text files.
Preprocessing
Each clinical note contains multiple medications, and we have to predict an event class for each medication mention. In the pre-processing, we created multiple sequences/text chunks for each medication mention in the form [context] medication [context]. A window based approach was used to determine the length of the context. After conducting multiple experiments a window-size of 200 characters achieved the best performance. However, in some cases this window size might not be sufficient to capture the context. The code used for pre-processing the dataset can be found here.
Models
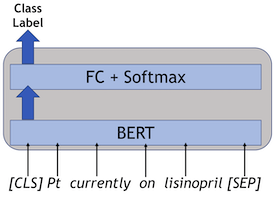
We approached this task as a sequence classification task and used the standard BERT model for sequence classification from huggingface. The best model was an ensemble of three BERT-based models, i.e., Clinical longformer, BioClinical BERT, and PubMed BERT. Since neural network models provide different results when initialized with different seeds, previous research suggests using different seeds and averaging the results. The code for the models can be found in this Github repository.
References
- https://n2c2.dbmi.hms.harvard.edu/2022-track-1
- Mahajan, D., Liang, J.J. and Tsou, C.H., 2020. Toward Understanding Clinical Context of Medication Change Events in Clinical Narratives. arXiv preprint arXiv:2011.08835
- Alsentzer E, Murphy J, Boag W, Weng W-H, Jindi D, Naumann T, et al. Publicly available Clinical BERT embeddings. In: Proceedings of the 2nd Clinical Natural Language Processing Workshop. Minneapolis, Minnesota, USA: ACL; 2019:72–8. [accessed 4 Jun 2020]
- Gu Y, Tinn R, Cheng H, Lucas M, Usuyama N, Liu X, Naumann T, Gao J, Poon H. Domain specific language model pretraining for biomedical natural language processing. ACM Transactions on Computing for Healthcare (HEALTH). 2021 Oct 15;3(1):1-23
- Li Y, Wehbe RM, Ahmad FS, Wang H, Luo Y. Clinical-Longformer and Clinical-BigBird: transformers for long clinical sequences. arXiv preprint arXiv:2201.11838. 2022 Jan 27